Downloading Data with rgbif
Preamble, Package-Loading, and GBIF API Credential Registering (click here):
## Custom install & load function
install.load.package <- function(x) {
if (!require(x, character.only = TRUE)) {
install.packages(x, repos = "http://cran.us.r-project.org")
}
require(x, character.only = TRUE)
}
## names of packages we want installed (if not installed yet) and loaded
package_vec <- c(
"rgbif",
"knitr", # for rmarkdown table visualisations
"rio" # for dwc import
)
## executing install & load for each package
sapply(package_vec, install.load.package)
## rgbif knitr rio
## TRUE TRUE TRUE
options(gbif_user = "my gbif username")
options(gbif_email = "my registred gbif e-mail")
options(gbif_pwd = "my gbif password")
First, we need to register the correct usageKey for Lagopus muta:
sp_name <- "Lagopus muta"
sp_backbone <- name_backbone(name = sp_name)
sp_key <- sp_backbone$usageKey
Second, this is the specific data we are interested in obtaining:
occ_final <- occ_search(
taxonKey = sp_key,
country = "NO",
year = "2000,2020",
basisOfRecord = "HUMAN_OBSERVATION",
occurrenceStatus = "PRESENT"
)
Finally, we are ready to obtain GBIF mediated data we may want to use in downstream analyses and applications. This can be done in one of two ways - either via:
- Near-Instant Download - via
occ_data(...) - Asynchronous Download - via
occ_download(...)
Let’s explore both of these in turn.
Near-Instant Download
Near-instant downloads do not require a GBIF account to be set-up and are the quickest way to obtain GBIF mediated data, but also the more limited functionality since near-instant downloads allow the user to only obtain 100,000 records per query and do not create citable DOIs for easy reference and accreditation of GBIF mediated data.
Let me demonstrate how these can be executed nevertheless.
As a matter of fact, we have already executed one of these through our call to occ_search(...) at the top of this page. While this function does download data via the ...$data output it provides, let’s showcase instead how we could use the occ_data(...) function to arrive at the same result:
NI_occ <- occ_data(
taxonKey = sp_key,
country = "NO",
year = "2000,2020",
basisOfRecord = "HUMAN_OBSERVATION",
occurrenceStatus = "PRESENT"
)
knitr::kable(head(NI_occ$data))
| key | scientificName | decimalLatitude | decimalLongitude | issues | datasetKey | publishingOrgKey | installationKey | hostingOrganizationKey | publishingCountry | protocol | lastCrawled | lastParsed | crawlId | basisOfRecord | individualCount | occurrenceStatus | taxonKey | kingdomKey | phylumKey | classKey | orderKey | familyKey | genusKey | speciesKey | acceptedTaxonKey | acceptedScientificName | kingdom | phylum | order | family | genus | species | genericName | specificEpithet | taxonRank | taxonomicStatus | iucnRedListCategory | continent | stateProvince | year | month | day | eventDate | startDayOfYear | endDayOfYear | lastInterpreted | license | isSequenced | isInCluster | recordedBy | geodeticDatum | class | countryCode | gbifRegion | country | publishedByGbifRegion | identifier | catalogNumber | vernacularName | institutionCode | taxonConceptID | locality | gbifID | collectionCode | occurrenceID | coordinateUncertaintyInMeters | modified | dynamicProperties | municipality | county | associatedReferences | locationID | fieldNotes | eventTime | behavior | sex | identificationVerificationStatus | identificationRemarks | dateIdentified | references | datasetName | identifiedBy | rightsHolder | http://unknown.org/nick | verbatimEventDate | verbatimLocality | taxonID | http://unknown.org/captive | identificationID | datasetID | footprintWKT | lifeStage | samplingProtocol | habitat | eventType | networkKeys | elevation | elevationAccuracy | organismQuantity | organismQuantityType | sampleSizeUnit | sampleSizeValue | eventID | informationWithheld | dataGeneralizations | samplingEffort | type | ownerInstitutionCode | occurrenceRemarks | name |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3219111975 | Lagopus muta (Montin, 1781) | 58.46553 | 8.895254 | cdc | 4fa7b334-ce0d-4e88-aaae-2e0c138d049e | e2e717bf-551a-4917-bdc9-4fa0f342c530 | 7182d304-b0a2-404b-baba-2086a325c221 | e2e717bf-551a-4917-bdc9-4fa0f342c530 | NO | DWC_ARCHIVE | 2024-04-15T22:36:22.136+00:00 | 2024-04-17T08:30:11.331+00:00 | 20 | HUMAN_OBSERVATION | 1 | PRESENT | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | 5227679 | 5227679 | Lagopus muta (Montin, 1781) | Animalia | Chordata | Galliformes | Phasianidae | Lagopus | Lagopus muta | Lagopus | muta | SPECIES | ACCEPTED | LC | EUROPE | Aust-Agder | 2020 | 1 | 5 | 2020-01-05 | 5 | 5 | 2024-04-17T08:30:11.331+00:00 | http://creativecommons.org/licenses/by/4.0/legalcode | FALSE | FALSE | obsr1460143 | WGS84 | Aves | NO | EUROPE | Norway | EUROPE | OBS854553845 | OBS854553845 | Rock Ptarmigan | CLO | avibase-79B161B7 | Botstangen | 3219111975 | EBIRD | URN:catalog:CLO:EBIRD:OBS854553845 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NULL | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | Lagopus muta (Montin, 1781) |
| 2560812506 | Lagopus muta (Montin, 1781) | 60.86029 | 8.513532 | cdc,cdround,gass84 | b124e1e0-4755-430f-9eab-894f25a9b59c | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | 7bdf9f6d-317a-45ec-8bb7-7ff61345d6a6 | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | NO | EML | 2024-11-12T15:07:16.815+00:00 | 2024-11-12T18:49:48.979+00:00 | 386 | HUMAN_OBSERVATION | 18 | PRESENT | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | 5227679 | 5227679 | Lagopus muta (Montin, 1781) | Animalia | Chordata | Galliformes | Phasianidae | Lagopus | Lagopus muta | Lagopus | muta | SPECIES | ACCEPTED | LC | EUROPE | Viken | 2020 | 1 | 26 | 2020-01-26 | 26 | 26 | 2024-11-12T18:49:48.979+00:00 | http://creativecommons.org/licenses/by/4.0/legalcode | FALSE | FALSE | Magnus Larsson | WGS84 | Aves | NO | EUROPE | Norway | EUROPE | urn:uuid:089a889f-e894-433c-a466-9e00708d212c | 23369915 | NA | nof | NA | Hemsedal Skisenter, Hemsedal, Vi | 2560812506 | so2-birds | urn:uuid:089a889f-e894-433c-a466-9e00708d212c | 300 | 2020-01-28T08:37:41.000+00:00 | {“Activity”:“Stationary”} | Hemsedal | Viken | https://www.artsobservasjoner.no/Sighting/23369915 | 394857 | Activity: Stationary. | 11.0/11.17 | stationary | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NULL | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | Lagopus muta (Montin, 1781) |
| 2544028549 | Lagopus muta (Montin, 1781) | 58.46407 | 8.895621 | cdc,cdround,gass84 | b124e1e0-4755-430f-9eab-894f25a9b59c | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | 7bdf9f6d-317a-45ec-8bb7-7ff61345d6a6 | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | NO | EML | 2024-11-12T15:07:16.815+00:00 | 2024-11-12T18:44:48.202+00:00 | 386 | HUMAN_OBSERVATION | 1 | PRESENT | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | 5227679 | 5227679 | Lagopus muta (Montin, 1781) | Animalia | Chordata | Galliformes | Phasianidae | Lagopus | Lagopus muta | Lagopus | muta | SPECIES | ACCEPTED | LC | EUROPE | Agder | 2020 | 1 | 5 | 2020-01-05 | 5 | 5 | 2024-11-12T18:44:48.202+00:00 | http://creativecommons.org/licenses/by/4.0/legalcode | FALSE | FALSE | Trond Nilsen | WGS84 | Aves | NO | EUROPE | Norway | EUROPE | urn:uuid:0bd9ed37-74f0-4bef-bd31-f9b6f45c5430 | 23266188 | NA | nof | NA | Blåmannen, Botne, Arendal, Ag | 2544028549 | so2-birds | urn:uuid:0bd9ed37-74f0-4bef-bd31-f9b6f45c5430 | 79 | 2020-01-05T17:57:17.000+00:00 | NA | Arendal | Agder | https://www.artsobservasjoner.no/Sighting/23266188 | 461661 | NA | 8.67/9.33 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NULL | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | Lagopus muta (Montin, 1781) |
| 2549610691 | Lagopus muta (Montin, 1781) | 59.49676 | 8.990860 | cdc,cdround,gass84 | b124e1e0-4755-430f-9eab-894f25a9b59c | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | 7bdf9f6d-317a-45ec-8bb7-7ff61345d6a6 | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | NO | EML | 2024-11-12T15:07:16.815+00:00 | 2024-11-12T18:45:33.729+00:00 | 386 | HUMAN_OBSERVATION | 6 | PRESENT | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | 5227679 | 5227679 | Lagopus muta (Montin, 1781) | Animalia | Chordata | Galliformes | Phasianidae | Lagopus | Lagopus muta | Lagopus | muta | SPECIES | ACCEPTED | LC | EUROPE | Vestfold og Telemark | 2020 | 1 | 10 | 2020-01-10 | 10 | 10 | 2024-11-12T18:45:33.729+00:00 | http://creativecommons.org/licenses/by/4.0/legalcode | FALSE | FALSE | Ola Nordsteien|Hanne Løvberg | WGS84 | Aves | NO | EUROPE | Norway | EUROPE | urn:uuid:25c5ecea-3187-4436-ab51-dc8438e78bb8 | 23288768 | NA | nof | NA | Skåråfjell, Lifjell, Midt-Telemark, Vt | 2549610691 | so2-birds | urn:uuid:25c5ecea-3187-4436-ab51-dc8438e78bb8 | 300 | 2020-01-10T23:48:45.000+00:00 | {“Activity”:“Forage”} | Midt-Telemark | Vestfold og Telemark | https://www.artsobservasjoner.no/Sighting/23288768 | 359743 | Activity: Forage. | NA | feeding | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NULL | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | Lagopus muta (Montin, 1781) |
| 2560817283 | Lagopus muta (Montin, 1781) | 69.77555 | 18.822903 | cdc,cdround,gass84 | b124e1e0-4755-430f-9eab-894f25a9b59c | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | 7bdf9f6d-317a-45ec-8bb7-7ff61345d6a6 | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | NO | EML | 2024-11-12T15:07:16.815+00:00 | 2024-11-12T18:50:14.238+00:00 | 386 | HUMAN_OBSERVATION | 1 | PRESENT | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | 5227679 | 5227679 | Lagopus muta (Montin, 1781) | Animalia | Chordata | Galliformes | Phasianidae | Lagopus | Lagopus muta | Lagopus | muta | SPECIES | ACCEPTED | LC | EUROPE | Troms og Finnmark | 2020 | 1 | 26 | 2020-01-26 | 26 | 26 | 2024-11-12T18:50:14.238+00:00 | http://creativecommons.org/licenses/by/4.0/legalcode | FALSE | FALSE | Ole-Morten Toften | WGS84 | Aves | NO | EUROPE | Norway | EUROPE | urn:uuid:2f576cfd-4387-4473-875c-b6ce9fa7113b | 23359018 | NA | nof | NA | Lyfjorddalen, Kvaløya, Tromsø, Tf | 2560817283 | so2-birds | urn:uuid:2f576cfd-4387-4473-875c-b6ce9fa7113b | 300 | 2020-01-26T13:07:00.000+00:00 | {“Activity”:“Display/SongOutsideBreeding”} | Tromsø | Troms og Finnmark | https://www.artsobservasjoner.no/Sighting/23359018 | 371536 | Activity: Display/SongOutsideBreeding. | 8.77 | stationary | MALE | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NULL | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | Lagopus muta (Montin, 1781) |
| 2544037194 | Lagopus muta (Montin, 1781) | 62.57669 | 11.387036 | cdc,cdround,gass84 | b124e1e0-4755-430f-9eab-894f25a9b59c | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | 7bdf9f6d-317a-45ec-8bb7-7ff61345d6a6 | d3978a37-635a-4ae3-bb85-7b4d41bc0b88 | NO | EML | 2024-11-12T15:07:16.815+00:00 | 2024-11-12T18:46:03.234+00:00 | 386 | HUMAN_OBSERVATION | 5 | PRESENT | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | 5227679 | 5227679 | Lagopus muta (Montin, 1781) | Animalia | Chordata | Galliformes | Phasianidae | Lagopus | Lagopus muta | Lagopus | muta | SPECIES | ACCEPTED | LC | EUROPE | Trøndelag | 2020 | 1 | 4 | 2020-01-04 | 4 | 4 | 2024-11-12T18:46:03.234+00:00 | http://creativecommons.org/licenses/by/4.0/legalcode | FALSE | FALSE | Norvald Grønning | WGS84 | Aves | NO | EUROPE | Norway | EUROPE | urn:uuid:3a154636-f513-46de-a3d6-6c2dfdeb3283 | 23267493 | NA | nof | NA | Røros by, Røros, Tø | 2544037194 | so2-birds | urn:uuid:3a154636-f513-46de-a3d6-6c2dfdeb3283 | 300 | 2021-09-22T11:26:11.000+00:00 | {“ValidationStatus”:“Approved Documented”} | Røros | Trøndelag | https://www.artsobservasjoner.no/Sighting/23267493 | 355523 | Validationstatus: Approved Documented | NA | NA | NA | validated | Validator: Andreas Winnem | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NULL | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | Lagopus muta (Montin, 1781) |
The near-instantaneous nature of this type of download is largely due to the hard limit on how much data you may obtain. For our example, the 100,000 record limit is not an issue as we are dealing with only 10606 records being available to us, but queries to GBIF mediated records can easily and quickly exceed this limit.
Asynchronous Download
To avoid the reproducibility and data richness limitations of near-instant downloads, we must make a more formal download query to the GBIF API and await processing of our data request - an asynchronous download. To do so, we need to do three things in order:
- Specify the data we want and request processing and download from GBIF
- Download the data once it is processed
- Load the downloaded data into
R
Let’s tackle these step-by-step.
Data Request at GBIF
To start this process of obtaining GBIF-mediated occurrence records, we first need to make a request to GBIF to start processing of the data we require. This is done with the occ_download(...) function. Making a data request at GBIF via the occ_download(...) function comes with two important considerations:
occ_download(...)specific syntax- data query metadata
Syntax and Query through occ_download()
The occ_download(...) function - while powerful - requires us to confront a new set of syntax which translates the download request as we have seen it so far into a form which the GBIF API can understand. To do so, we use a host of rgbif functions built around the GBIF predicate DSL (domain specific language). These functions (with a few exceptions which you can see by calling the documentation - ?download_predicate_dsl) all take two arguments:
key- this is a core term which we want to target for our requestvalue- this is the value for the core term which we are interested in
Finally, the relevant functions and how the relate key to value are:
pred(...): equalspred_lt(...): lessThanpred_lte(...): lessThanOrEqualspred_gt(...): greaterThanpred_gte(...): greaterThanOrEqualspred_like(...): likepred_within(...): within (only for geospatial queries, and only accepts a WKT string)pred_notnull(...): isNotNull (only for geospatial queries, and only accepts a WKT string)pred_isnull(...): isNull (only for stating that you want a key to be null)pred_and(...): and (accepts multiple individual predicates, separating them by either “and” or “or”)pred_or(...): or (accepts multiple individual predicates, separating them by either “and” or “or”)pred_not(...): not (negates whatever predicate is passed in)pred_in(...): in (accepts a single key but many values; stating that you want to search for all the values)
Let us use these to query the data we are interested in:
res <- occ_download(
pred("taxonKey", sp_key),
pred("basisOfRecord", "HUMAN_OBSERVATION"),
pred("country", "NO"),
pred("hasCoordinate", TRUE),
pred_gte("year", 2000),
pred_lte("year", 2020)
)
Now that the download request has been made with GBIF, you need to wait for GBIF to process your query and return the desired data to you. You will find an overview of all your requested and processed downloads in your personal download tab on the GBIF webportal.
Data Query Metadata
To keep track of all your data queries, each call to occ_download(...) returns a set of metadata for your download request as part of its object structure. This can be assessed as follows:
occ_download_meta(res)
## <<gbif download metadata>>
## Status: SUCCEEDED
## DOI: 10.15468/dl.vjazpv
## Format: DWCA
## Download key: 0010612-241024112534372
## Created: 2024-10-29T13:25:46.576+00:00
## Modified: 2024-10-29T13:27:15.574+00:00
## Download link: https://api.gbif.org/v1/occurrence/download/request/0010612-241024112534372.zip
## Total records: 10609
It can even be subset for relevant parts of it like so:
occ_download_meta(res)$downloadLink
## [1] "https://api.gbif.org/v1/occurrence/download/request/0010612-241024112534372.zip"
This is done according to the underlying structure of the metadata:
str(occ_download_meta(res), max.level = 1)
## List of 12
## $ key : chr "0010612-241024112534372"
## $ doi : chr "10.15468/dl.vjazpv"
## $ license : chr "http://creativecommons.org/licenses/by-nc/4.0/legalcode"
## $ request :List of 5
## $ created : chr "2024-10-29T13:25:46.576+00:00"
## $ modified : chr "2024-10-29T13:27:15.574+00:00"
## $ eraseAfter : chr "2025-04-29T13:25:46.526+00:00"
## $ status : chr "SUCCEEDED"
## $ downloadLink : chr "https://api.gbif.org/v1/occurrence/download/request/0010612-241024112534372.zip"
## $ size : int 2842371
## $ totalRecords : int 10609
## $ numberDatasets: int 18
## - attr(*, "class")= chr "occ_download_meta"
## - attr(*, "format")= chr "DWCA"
Note: You will find that the number of records in our query made via occ_download(...) here and occ_search(...) above are different. This is due to different ways data ranges are handled.
occ_download_meta(res)$totalRecords
## [1] 10609
Data Download
Now that the download request is registered with GBIF, it is time to actually obtain the data you have queried. To facilitate this, there are two distinct ways to obtain the data itself:
- Webportal
rgbif
Personally, I can not recommend any of these methods over the rgbif-internal download method. However, in case you might prefer other options, I will quickly show how these work, too.
GBIF Portal Download Method
At the most basic level, you may obtain data through the GBIF portal where you will find an overview of your download requests. Any individual download request can be investigated more deeply by clicking the “SHOW” button:
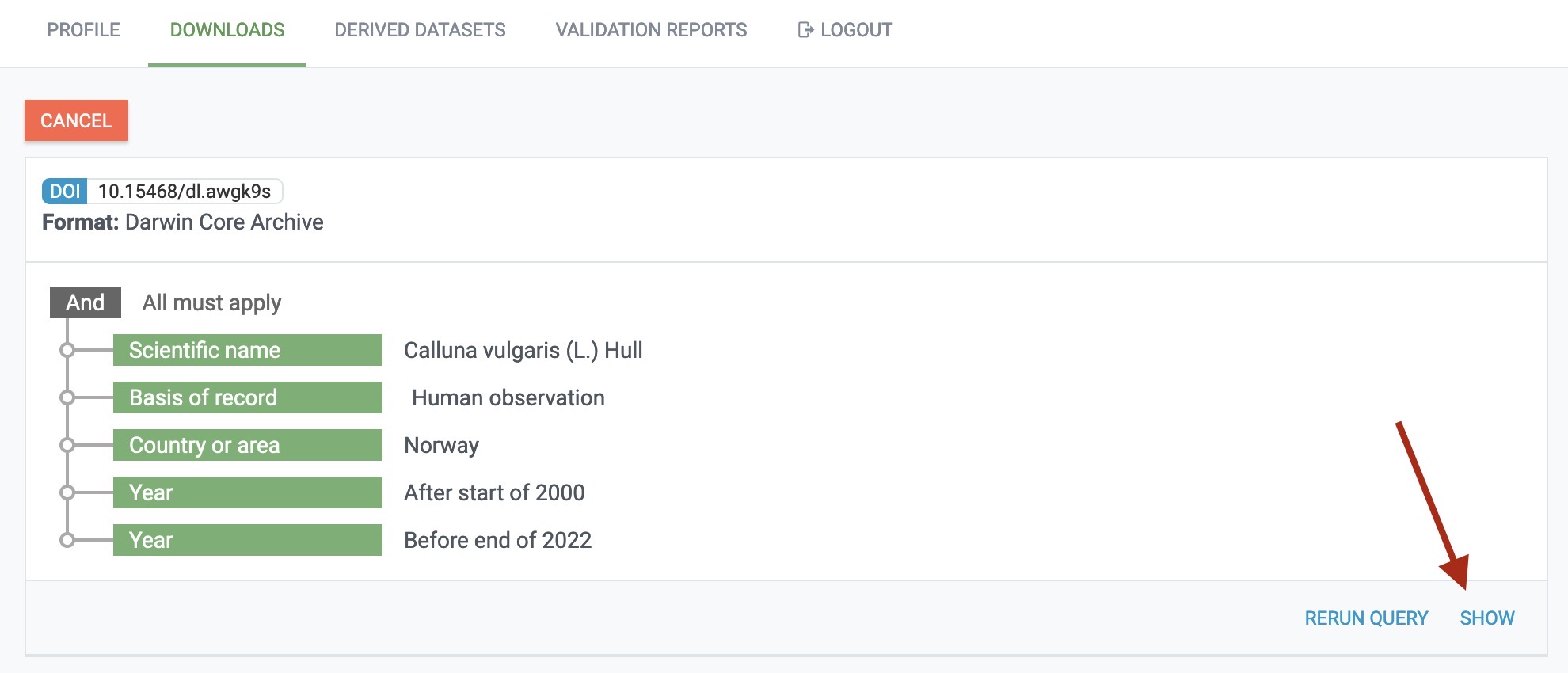
There, you will see a breakdown of your download request and also whether it is still being processed or already finished:
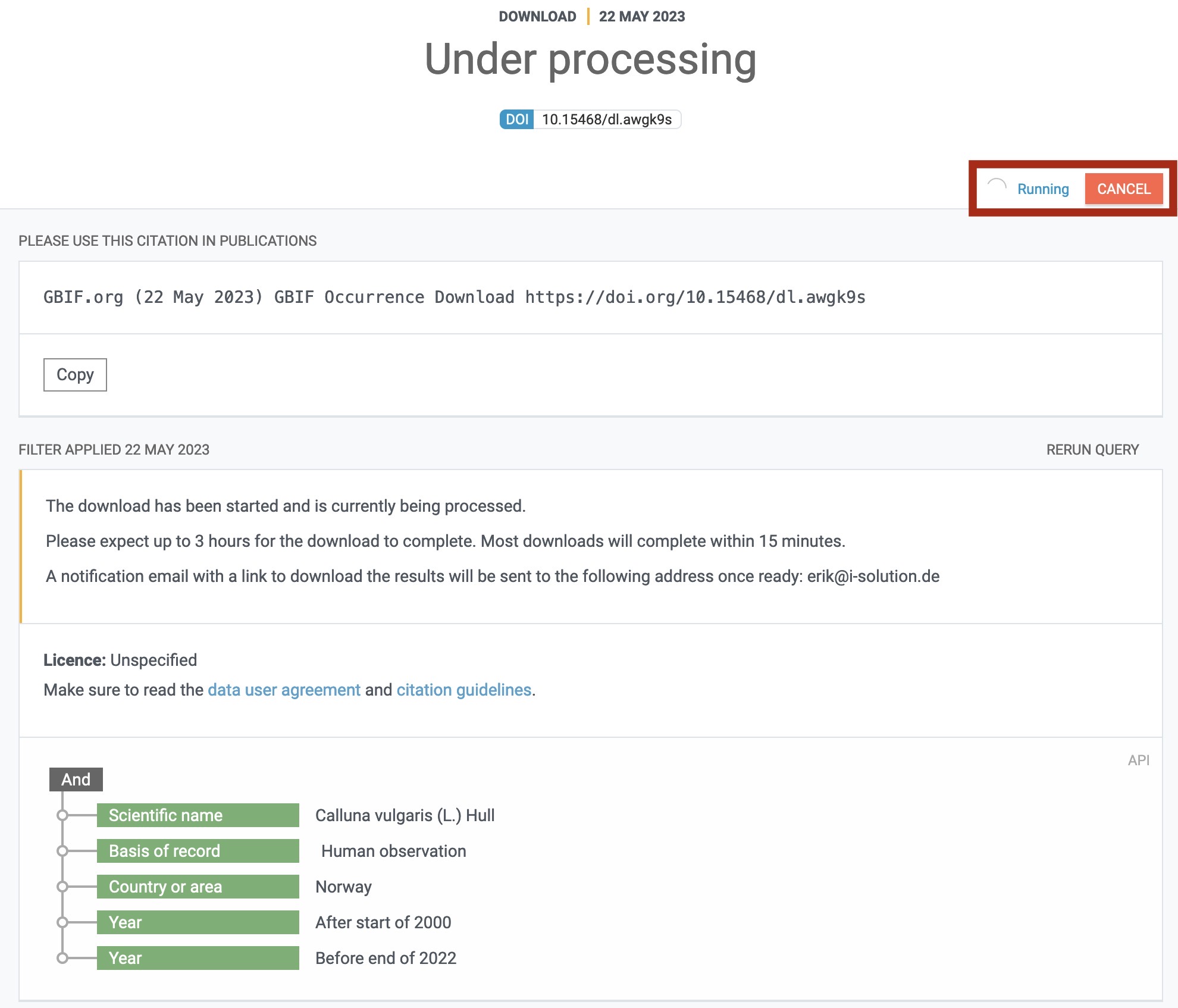
Note that the DOI differs here from the other images and the data product we are working with ultimately. This is because I had to requeue the download to get this screenshot.
Upon completion of your request on the GBIF end, a “DOWNLOAD” button will appear on the specific download request overview page:
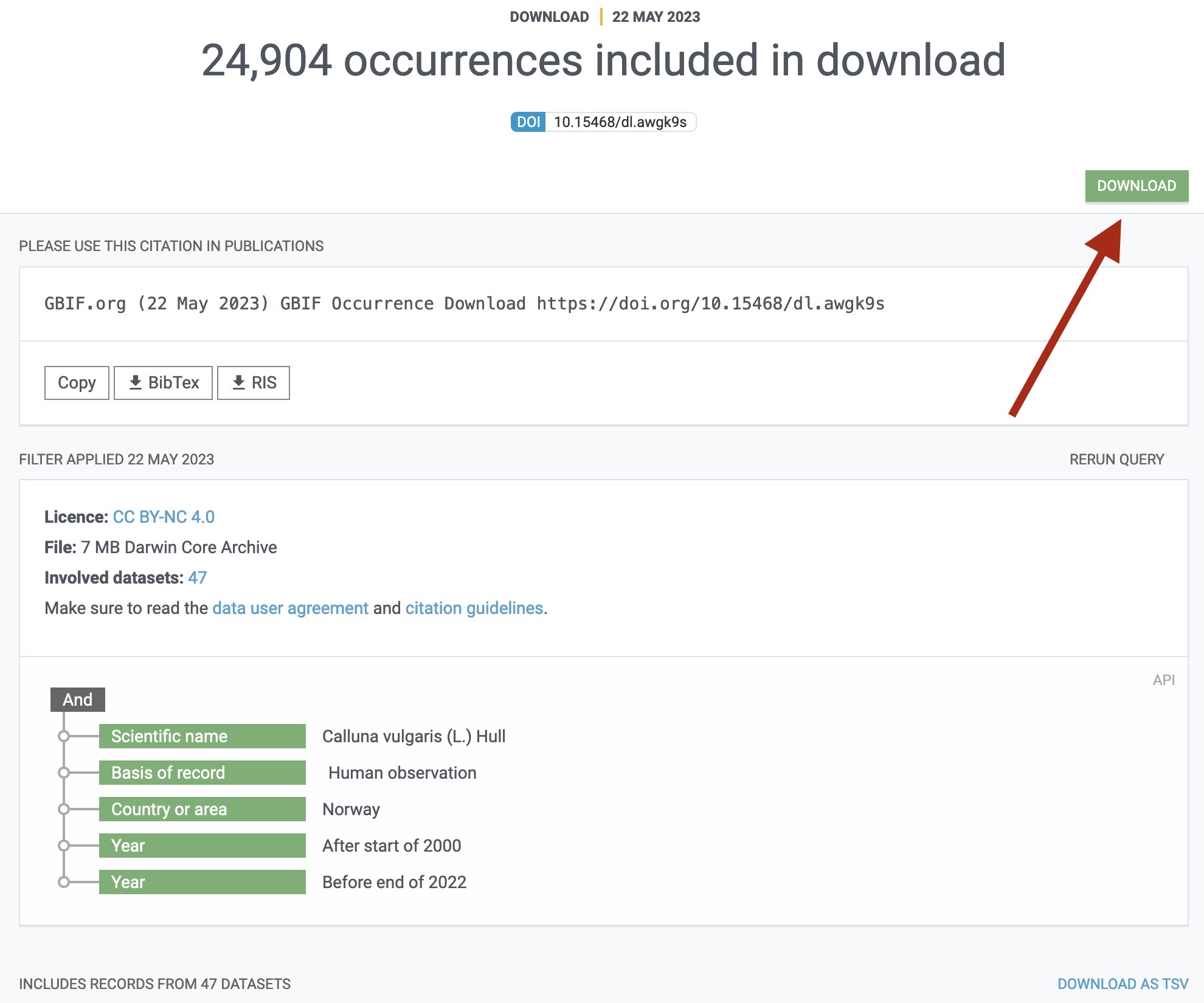
For requests of few data points you will usually have to be quite quick to see this in action as GBIF is often pretty fast and getting data ready for you.
rgbif Download Method
To download our data programmatically, we need to execute roughly the same steps as we did above:
- Wait for data to be ready for download
- Download the data
- Load the data into
R(we didn’t do this in the above)
This is done with the following three corresponding functions:
## 1. Check GBIF whether data is ready, this function will finish running when done and return metadata
res_meta <- occ_download_wait(res, status_ping = 5, curlopts = list(), quiet = FALSE)
## 2. Download the data as .zip (can specify a path)
res_get <- occ_download_get(res)
## 3. Load the data into R
res_data <- occ_download_import(res_get)
Let’s have a look at the data we just loaded:
dim(res_data)
## [1] 10609 223
knitr::kable(head(res_data))
| gbifID | accessRights | bibliographicCitation | language | license | modified | publisher | references | rightsHolder | type | institutionID | collectionID | datasetID | institutionCode | collectionCode | datasetName | ownerInstitutionCode | basisOfRecord | informationWithheld | dataGeneralizations | dynamicProperties | occurrenceID | catalogNumber | recordNumber | recordedBy | recordedByID | individualCount | organismQuantity | organismQuantityType | sex | lifeStage | reproductiveCondition | caste | behavior | vitality | establishmentMeans | degreeOfEstablishment | pathway | georeferenceVerificationStatus | occurrenceStatus | preparations | disposition | associatedOccurrences | associatedReferences | associatedSequences | associatedTaxa | otherCatalogNumbers | occurrenceRemarks | organismID | organismName | organismScope | associatedOrganisms | previousIdentifications | organismRemarks | materialEntityID | materialEntityRemarks | verbatimLabel | materialSampleID | eventID | parentEventID | eventType | fieldNumber | eventDate | eventTime | startDayOfYear | endDayOfYear | year | month | day | verbatimEventDate | habitat | samplingProtocol | sampleSizeValue | sampleSizeUnit | samplingEffort | fieldNotes | eventRemarks | locationID | higherGeographyID | higherGeography | continent | waterBody | islandGroup | island | countryCode | stateProvince | county | municipality | locality | verbatimLocality | verbatimElevation | verticalDatum | verbatimDepth | minimumDistanceAboveSurfaceInMeters | maximumDistanceAboveSurfaceInMeters | locationAccordingTo | locationRemarks | decimalLatitude | decimalLongitude | coordinateUncertaintyInMeters | coordinatePrecision | pointRadiusSpatialFit | verbatimCoordinateSystem | verbatimSRS | footprintWKT | footprintSRS | footprintSpatialFit | georeferencedBy | georeferencedDate | georeferenceProtocol | georeferenceSources | georeferenceRemarks | geologicalContextID | earliestEonOrLowestEonothem | latestEonOrHighestEonothem | earliestEraOrLowestErathem | latestEraOrHighestErathem | earliestPeriodOrLowestSystem | latestPeriodOrHighestSystem | earliestEpochOrLowestSeries | latestEpochOrHighestSeries | earliestAgeOrLowestStage | latestAgeOrHighestStage | lowestBiostratigraphicZone | highestBiostratigraphicZone | lithostratigraphicTerms | group | formation | member | bed | identificationID | verbatimIdentification | identificationQualifier | typeStatus | identifiedBy | identifiedByID | dateIdentified | identificationReferences | identificationVerificationStatus | identificationRemarks | taxonID | scientificNameID | acceptedNameUsageID | parentNameUsageID | originalNameUsageID | nameAccordingToID | namePublishedInID | taxonConceptID | scientificName | acceptedNameUsage | parentNameUsage | originalNameUsage | nameAccordingTo | namePublishedIn | namePublishedInYear | higherClassification | kingdom | phylum | class | order | superfamily | family | subfamily | tribe | subtribe | genus | genericName | subgenus | infragenericEpithet | specificEpithet | infraspecificEpithet | cultivarEpithet | taxonRank | verbatimTaxonRank | vernacularName | nomenclaturalCode | taxonomicStatus | nomenclaturalStatus | taxonRemarks | datasetKey | publishingCountry | lastInterpreted | elevation | elevationAccuracy | depth | depthAccuracy | distanceFromCentroidInMeters | issue | mediaType | hasCoordinate | hasGeospatialIssues | taxonKey | acceptedTaxonKey | kingdomKey | phylumKey | classKey | orderKey | familyKey | genusKey | subgenusKey | speciesKey | species | acceptedScientificName | verbatimScientificName | typifiedName | protocol | lastParsed | lastCrawled | repatriated | relativeOrganismQuantity | projectId | isSequenced | gbifRegion | publishedByGbifRegion | level0Gid | level0Name | level1Gid | level1Name | level2Gid | level2Name | level3Gid | level3Name | iucnRedListCategory |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 979205774 | NA | NA | CC_BY_4_0 | NA | NA | NA | NA | NA | CLO | EBIRD | HUMAN_OBSERVATION | URN:catalog:CLO:EBIRD:OBS238252603 | OBS238252603 | NA | obsr498576 | NA | 1 | NA | NA | NA | NA | NA | NA | NA | NA | PRESENT | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | 2011-05-12 | 132 | 132 | 2011 | 5 | 12 | NA | NA | EUROPE | NA | NA | NA | NO | Oppland | Fisketjerni | NA | NA | NA | NA | NA | 61.37563 | 8.840089 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | avibase-79B161B7 | Lagopus muta (Montin, 1781) | NA | NA | NA | NA | NA | Animalia | Chordata | Aves | Galliformes | NA | Phasianidae | NA | NA | NA | Lagopus | Lagopus | NA | NA | muta | NA | SPECIES | NA | Rock Ptarmigan | ACCEPTED | NA | NA | 4fa7b334-ce0d-4e88-aaae-2e0c138d049e | NO | 2024-04-17T08:24:53.730Z | NA | NA | NA | NA | NA | CONTINENT_DERIVED_FROM_COORDINATES;TAXON_MATCH_TAXON_CONCEPT_ID_IGNORED | TRUE | FALSE | 5227679 | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | NA | 5227679 | Lagopus muta | Lagopus muta (Montin, 1781) | Lagopus muta | NA | DWC_ARCHIVE | 2024-04-17T08:24:53.730Z | 2024-04-15T22:36:22.136Z | FALSE | NA | FALSE | EUROPE | EUROPE | NOR | Norway | NOR.11_1 | Oppland | NOR.11.16_1 | Øystre Slidre | NA | NA | LC | |||||||||||||||||||||||||||||||||||||||||||||||||||||
| 977864468 | NA | NA | CC_BY_4_0 | NA | NA | NA | NA | NA | CLO | EBIRD | HUMAN_OBSERVATION | URN:catalog:CLO:EBIRD:OBS227313790 | OBS227313790 | NA | obsr419757 | NA | 2 | NA | NA | NA | NA | NA | NA | NA | NA | PRESENT | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | 2007-06-09 | 160 | 160 | 2007 | 6 | 9 | NA | NA | EUROPE | NA | NA | NA | NO | Finnmark | Syltefjord | NA | NA | NA | NA | NA | 70.54530 | 30.054518 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | avibase-79B161B7 | Lagopus muta (Montin, 1781) | NA | NA | NA | NA | NA | Animalia | Chordata | Aves | Galliformes | NA | Phasianidae | NA | NA | NA | Lagopus | Lagopus | NA | NA | muta | NA | SPECIES | NA | Rock Ptarmigan | ACCEPTED | NA | NA | 4fa7b334-ce0d-4e88-aaae-2e0c138d049e | NO | 2024-04-17T08:23:18.063Z | NA | NA | NA | NA | NA | CONTINENT_DERIVED_FROM_COORDINATES;TAXON_MATCH_TAXON_CONCEPT_ID_IGNORED | TRUE | FALSE | 5227679 | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | NA | 5227679 | Lagopus muta | Lagopus muta (Montin, 1781) | Lagopus muta | NA | DWC_ARCHIVE | 2024-04-17T08:23:18.063Z | 2024-04-15T22:36:22.136Z | FALSE | NA | FALSE | EUROPE | EUROPE | NOR | Norway | NOR.5_1 | Finnmark | NOR.5.2_1 | Båtsfjord | NA | NA | LC | |||||||||||||||||||||||||||||||||||||||||||||||||||||
| 977848277 | NA | NA | CC_BY_4_0 | NA | NA | NA | NA | NA | CLO | EBIRD | HUMAN_OBSERVATION | URN:catalog:CLO:EBIRD:OBS227314225 | OBS227314225 | NA | obsr419757 | NA | 1 | NA | NA | NA | NA | NA | NA | NA | NA | PRESENT | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | 2007-06-09 | 160 | 160 | 2007 | 6 | 9 | NA | NA | EUROPE | NA | NA | NA | NO | Finnmark | Julahaugen | NA | NA | NA | NA | NA | 70.52452 | 29.125786 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | avibase-79B161B7 | Lagopus muta (Montin, 1781) | NA | NA | NA | NA | NA | Animalia | Chordata | Aves | Galliformes | NA | Phasianidae | NA | NA | NA | Lagopus | Lagopus | NA | NA | muta | NA | SPECIES | NA | Rock Ptarmigan | ACCEPTED | NA | NA | 4fa7b334-ce0d-4e88-aaae-2e0c138d049e | NO | 2024-04-17T08:23:56.208Z | NA | NA | NA | NA | NA | CONTINENT_DERIVED_FROM_COORDINATES;TAXON_MATCH_TAXON_CONCEPT_ID_IGNORED | TRUE | FALSE | 5227679 | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | NA | 5227679 | Lagopus muta | Lagopus muta (Montin, 1781) | Lagopus muta | NA | DWC_ARCHIVE | 2024-04-17T08:23:56.208Z | 2024-04-15T22:36:22.136Z | FALSE | NA | FALSE | EUROPE | EUROPE | NOR | Norway | NOR.5_1 | Finnmark | NOR.5.3_1 | Berlevåg | NA | NA | LC | |||||||||||||||||||||||||||||||||||||||||||||||||||||
| 976560970 | NA | NA | CC_BY_4_0 | NA | NA | NA | NA | NA | CLO | EBIRD | HUMAN_OBSERVATION | URN:catalog:CLO:EBIRD:OBS223362772 | OBS223362772 | NA | obsr226921 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | PRESENT | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | 2010-05-29 | 149 | 149 | 2010 | 5 | 29 | NA | NA | EUROPE | NA | NA | NA | NO | Sør-Trøndelag | Norway, Kongsvold | NA | NA | NA | NA | NA | 62.30637 | 9.609218 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | avibase-79B161B7 | Lagopus muta (Montin, 1781) | NA | NA | NA | NA | NA | Animalia | Chordata | Aves | Galliformes | NA | Phasianidae | NA | NA | NA | Lagopus | Lagopus | NA | NA | muta | NA | SPECIES | NA | Rock Ptarmigan | ACCEPTED | NA | NA | 4fa7b334-ce0d-4e88-aaae-2e0c138d049e | NO | 2024-04-17T09:50:27.030Z | NA | NA | NA | NA | NA | CONTINENT_DERIVED_FROM_COORDINATES;TAXON_MATCH_TAXON_CONCEPT_ID_IGNORED | TRUE | FALSE | 5227679 | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | NA | 5227679 | Lagopus muta | Lagopus muta (Montin, 1781) | Lagopus muta | NA | DWC_ARCHIVE | 2024-04-17T09:50:27.030Z | 2024-04-15T22:36:22.136Z | FALSE | NA | FALSE | EUROPE | EUROPE | NOR | Norway | NOR.15_1 | Sør-Trøndelag | NOR.15.13_1 | Oppdal | NA | NA | LC | |||||||||||||||||||||||||||||||||||||||||||||||||||||
| 956725853 | NA | NA | CC_BY_4_0 | NA | NA | NA | NA | NA | CLO | EBIRD | HUMAN_OBSERVATION | URN:catalog:CLO:EBIRD:OBS201175760 | OBS201175760 | NA | obsr98173 | NA | 2 | NA | NA | NA | NA | NA | NA | NA | NA | PRESENT | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | 2013-06-12 | 163 | 163 | 2013 | 6 | 12 | NA | NA | EUROPE | NA | NA | NA | NO | Nordland | Arctic Circle - E6 | NA | NA | NA | NA | NA | 66.55537 | 15.315628 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | avibase-79B161B7 | Lagopus muta (Montin, 1781) | NA | NA | NA | NA | NA | Animalia | Chordata | Aves | Galliformes | NA | Phasianidae | NA | NA | NA | Lagopus | Lagopus | NA | NA | muta | NA | SPECIES | NA | Rock Ptarmigan | ACCEPTED | NA | NA | 4fa7b334-ce0d-4e88-aaae-2e0c138d049e | NO | 2024-04-17T08:23:05.601Z | NA | NA | NA | NA | NA | CONTINENT_DERIVED_FROM_COORDINATES;TAXON_MATCH_TAXON_CONCEPT_ID_IGNORED | TRUE | FALSE | 5227679 | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | NA | 5227679 | Lagopus muta | Lagopus muta (Montin, 1781) | Lagopus muta | NA | DWC_ARCHIVE | 2024-04-17T08:23:05.601Z | 2024-04-15T22:36:22.136Z | FALSE | NA | FALSE | EUROPE | EUROPE | NOR | Norway | NOR.10_1 | Nordland | NOR.10.28_1 | Rana | NA | NA | LC | |||||||||||||||||||||||||||||||||||||||||||||||||||||
| 922124083 | NA | NA | CC_BY_4_0 | NA | NA | NA | NA | NA | naturgucker | naturgucker | HUMAN_OBSERVATION | 1450641070 | NA | -1963296749 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | PRESENT | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | 2011-08-04T00:00 | 216 | 216 | 2011 | 8 | 4 | NA | NA | EUROPE | NA | NA | NA | NO | Ferienhaus Brekkeseter | NA | NA | NA | NA | NA | 61.89373 | 9.020678 | 250 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | Lagopus muta (Montin, 1781) | NA | NA | NA | NA | NA | Animalia | Chordata | Aves | Galliformes | NA | Phasianidae | NA | NA | NA | Lagopus | Lagopus | NA | NA | muta | NA | SPECIES | NA | ACCEPTED | NA | NA | 6ac3f774-d9fb-4796-b3e9-92bf6c81c084 | DE | 2024-03-15T23:26:53.443Z | NA | NA | NA | NA | NA | COORDINATE_ROUNDED;GEODETIC_DATUM_ASSUMED_WGS84;CONTINENT_DERIVED_FROM_COORDINATES;MULTIMEDIA_URI_INVALID | TRUE | FALSE | 5227679 | 5227679 | 1 | 44 | 212 | 723 | 9331 | 2473369 | NA | 5227679 | Lagopus muta | Lagopus muta (Montin, 1781) | Lagopus muta | NA | BIOCASE | 2024-03-15T23:26:53.443Z | 2024-03-15T21:43:29.197Z | TRUE | NA | FALSE | EUROPE | EUROPE | NOR | Norway | NOR.11_1 | Oppland | NOR.11.23_1 | Vågå | NA | NA | LC |
Note that the structure of this data set is different to that obtained with occ_search(...) hence enforcing that you should always stage downloads whose data are used in publications with occ_download(...)!
Session Info
## R version 4.4.0 (2024-04-24 ucrt)
## Platform: x86_64-w64-mingw32/x64
## Running under: Windows 11 x64 (build 22631)
##
## Matrix products: default
##
##
## locale:
## [1] LC_COLLATE=Norwegian Bokmål_Norway.utf8 LC_CTYPE=Norwegian Bokmål_Norway.utf8
## [3] LC_MONETARY=Norwegian Bokmål_Norway.utf8 LC_NUMERIC=C
## [5] LC_TIME=Norwegian Bokmål_Norway.utf8
##
## time zone: Europe/Oslo
## tzcode source: internal
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] rio_1.2.3 knitr_1.48 rgbif_3.8.1
##
## loaded via a namespace (and not attached):
## [1] styler_1.10.3 sass_0.4.9 utf8_1.2.4 generics_0.1.3
## [5] xml2_1.3.6 blogdown_1.19 stringi_1.8.4 httpcode_0.3.0
## [9] digest_0.6.37 magrittr_2.0.3 evaluate_0.24.0 grid_4.4.0
## [13] bookdown_0.40 fastmap_1.2.0 R.oo_1.26.0 R.cache_0.16.0
## [17] plyr_1.8.9 jsonlite_1.8.8 R.utils_2.12.3 whisker_0.4.1
## [21] crul_1.5.0 urltools_1.7.3 httr_1.4.7 purrr_1.0.2
## [25] fansi_1.0.6 scales_1.3.0 oai_0.4.0 lazyeval_0.2.2
## [29] jquerylib_0.1.4 cli_3.6.3 rlang_1.1.4 triebeard_0.4.1
## [33] R.methodsS3_1.8.2 bit64_4.0.5 munsell_0.5.1 cachem_1.1.0
## [37] yaml_2.3.10 tools_4.4.0 dplyr_1.1.4 colorspace_2.1-1
## [41] ggplot2_3.5.1 curl_5.2.2 vctrs_0.6.5 R6_2.5.1
## [45] lifecycle_1.0.4 stringr_1.5.1 bit_4.0.5 pkgconfig_2.0.3
## [49] pillar_1.9.0 bslib_0.8.0 gtable_0.3.6 data.table_1.16.0
## [53] glue_1.7.0 Rcpp_1.0.13 xfun_0.47 tibble_3.2.1
## [57] tidyselect_1.2.1 htmltools_0.5.8.1 rmarkdown_2.28 compiler_4.4.0